Simples, and no need for physical shearing methods. 
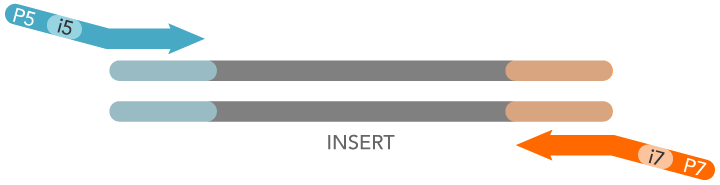
With an additional round of PCR, sequencing adaptors and multiplexing barcodes are incorporated into the fragment ends. Nextera is a really useful technique it uses a transpososome (a transposase and transposon end complex) to fragment the genome (semi-)randomly with the addition of a specific sequence. This is due to the introduction of Nextera library preparations. However, a recent analysis of a troublesome dataset has led me to revise my thoughts on the need to trim sequences routinely. Increasingly I found that quality was not a major determinant for getting good quality de novo assemblies, rather it came down to the usual old chestnuts of read length, coverage depth and insert size (for paired-end sequencing). Nowadays with the MiSeq and HiSeq instruments, read qualities still decline at the 3' end, but mainly remain usable. At the time I was dealing with early GA2 data, which had serious 3' quality drop-off issues, even with reads as short as 36 cycles (that's the true definition of short-read sequencing!). The explanation was likely due to the evolution of Illumina base-calling accuracy. I was increasingly finding that aggressive quality trimming was making little to no difference to my de novo assemblies, and in some cases actually making them worse. To date, it has been viewed a whopping 8,560 times!īut funnily enough, since that post was written my attitude towards filtering Illumina data slowly changed. One of the first posts I did on this blog, way back in September 2009 was about my experiences with filtering and trimming Illumina sequences, and it proved rather popular. The first record contains the right-caret character followed by an arbitrary string. The second record contains the adapter sequence. This file can contain multiple adapter sequences by using a multi-FASTA file format. Trimmomatic output files will show which reads (if any) were trimmed.Adaptor trim or die: Experiences with Nextera libraries In our example, using the Nextera XT library prep kit, the “adapters.fasta” file would look like this:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
